Playground S6E2 PRED: CV-Tuned Boosting Pipeline
Predicting Heart Disease: CV-Tuned Boosting (S6E2)
Kaggle notebook link: https://www.kaggle.com/code/pilkwang/s6e2-cv-tuned-boosting
The key idea is simple: keep the evaluation protocol fixed, then compare model families and data-augmentation variants fairly.
TL;DR
- Core objective: maximize reliable generalization, not one-off fold luck
- Validation:
StratifiedKFold(5)+ multi-seed averaging ([42, 2024, 7, 3407, 777]) - Main metric:
OOF ROC-AUC - Best run in notebook output:
CatBoost (base)withcv_auc = 0.955269 +originalaugmentation was competitive but not better thanbasein final ranking- Final submission generated from the top leaderboard row
- Public leaderboard score:
0.95334
Step 0. Imports and Global Config
Purpose
This section exists to make the rest of the notebook reproducible and comparable. Without fixed seeds and a shared global config, model-vs-model comparison is noisy and often misleading.
Parameter choices and why
SEED = 42,N_SPLITS = 5: standard reproducible CV baseline.SEED_LIST = [42, 2024, 7, 3407, 777]: multi-seed averaging to reduce leaderboard shake-up.TARGET_MAP = {'Absence': 0, 'Presence': 1}: unify label handling across train/original.CATS,NUMS: same feature typing as EDA to keep the pipeline consistent.ORIGINAL_WEIGHT = 0.35: default mixing weight for optional external data.
Outcome in this notebook
The config supported consistent model comparison across all candidate models and variants.
We evaluate each model on OOF ROC-AUC:
\[\mathrm{AUC} = \int_0^1 \mathrm{TPR}(u)\, d(\mathrm{FPR}(u))\]Quick notes for beginners:
StratifiedKFold: splits data into folds while preserving class ratio in each fold.roc_auc_score: computes threshold-independent ranking quality (AUC).TARGET_MAP: converts string labels into model-friendly numeric labels.
Show key code snippet
Reduced from original notebook: plotting style and non-essential logging lines were omitted.
1
2
3
4
5
6
7
8
9
10
11
12
13
14
SEED = 42
N_SPLITS = 5
SEED_LIST = [42, 2024, 7, 3407, 777]
TARGET = 'Heart Disease'
TARGET_BIN = 'target_bin'
ID_COL = 'id'
TARGET_MAP = {'Absence': 0, 'Presence': 1}
CATS = ['Sex', 'Chest pain type', 'FBS over 120', 'EKG results',
'Exercise angina', 'Slope of ST', 'Number of vessels fluro', 'Thallium']
NUMS = ['Age', 'BP', 'Cholesterol', 'Max HR', 'ST depression']
ORIGINAL_WEIGHT = 0.35
Step 1. Load Data
Purpose
This step confirms the training/inference schema and checks whether optional external data can be used for augmentation.
Parameter choices and why
- Load three competition files (
train,test,sample_submission) plus optionaloriginal. - Assert target/id schema early to fail fast if input is broken.
Outcome in this notebook
train:(630000, 15)test:(270000, 14)sample_submission:(270000, 2)original(optional):(270, 14)loaded successfully
This confirmed that augmentation experiments were available.
Quick notes for beginners:
assert ...: a fast sanity check that stops execution if schema is broken.HAS_ORIGINAL: runtime flag to enable/disable augmentation experiments automatically.
Show key code snippet
Reduced from original notebook: memory-print formatting lines were omitted.
1
2
3
4
5
6
7
8
9
10
11
12
13
train = pd.read_csv('/kaggle/input/playground-series-s6e2/train.csv')
test = pd.read_csv('/kaggle/input/playground-series-s6e2/test.csv')
sample_sub = pd.read_csv('/kaggle/input/playground-series-s6e2/sample_submission.csv')
try:
original = pd.read_csv('/kaggle/input/datasets/pilkwang/heart-disease-prediction/Heart_Disease_Prediction.csv')
HAS_ORIGINAL = True
except FileNotFoundError:
original = None
HAS_ORIGINAL = False
assert TARGET in train.columns
assert ID_COL in train.columns and ID_COL in test.columns
Step 2. Preprocessing (EDA-Aligned)
Purpose
Align feature/label preprocessing with EDA assumptions so model training does not drift from the statistical interpretation phase.
Parameter choices and why
- Build predictors by excluding
[id, target, target_bin]. - Map labels to binary once, then train all models on the same target definition.
- Unify categorical levels across
train/test/originalto prevent fold-time encoding mismatch.
Outcome in this notebook
This created stable X, y, X_test, and optional X_orig, y_orig inputs and prevented category-code inconsistency during CV.
Quick notes for beginners:
pd.CategoricalDtype(categories=...): forces train/test to share the same category universe.- This prevents fold-time or inference-time code mismatches for categorical models.
Show key code snippet
Reduced from original notebook: verbose diagnostics were omitted.
1
2
3
4
5
6
7
8
9
10
11
12
13
14
15
16
17
18
19
feature_cols = [c for c in train.columns if c not in [ID_COL, TARGET, TARGET_BIN]]
cats = [c for c in CATS if c in feature_cols]
nums = [c for c in NUMS if c in feature_cols]
train[TARGET_BIN] = train[TARGET].map(TARGET_MAP)
X = train[feature_cols].copy()
y = train[TARGET_BIN].astype('int8').copy()
X_test = test[feature_cols].copy()
if HAS_ORIGINAL:
original[TARGET_BIN] = original[TARGET].map(TARGET_MAP)
X_orig = original[feature_cols].copy()
y_orig = original[TARGET_BIN].astype('int8').copy()
for c in cats:
unified = pd.Index(sorted(set().union(*[set(df[c].dropna().unique()) for df in [X, X_test] + ([X_orig] if HAS_ORIGINAL else [])])))
cat_dtype = pd.CategoricalDtype(categories=unified)
for df in [X, X_test] + ([X_orig] if HAS_ORIGINAL else []):
df[c] = df[c].astype(cat_dtype)
Step 2.1 Optional Interaction Features
Purpose
Translate EDA interaction hints into concrete engineered features and test whether they help ranking quality.
Parameter choices and why
USE_INTERACTIONS = Truein this notebook run.- Numeric interactions: multiplicative and differential transforms (
Age x ST,MaxHR - Age, etc.). - Categorical interactions: pair-token columns for known high-signal combinations.
Outcome in this notebook
Feature space was expanded with interaction terms before model comparison. Final leaderboard shows competitive performance across all boosted models under this setup.
Quick notes for beginners:
- Interaction feature: a new feature built from two existing features (e.g., product or pair token).
astype('float32'): reduces memory while keeping enough precision for most tabular models.
Show key code snippet
Reduced from original notebook: memory-optimization lines were omitted.
1
2
3
4
5
6
7
8
9
10
11
12
13
14
USE_INTERACTIONS = True
if USE_INTERACTIONS:
for df in [X, X_test] + ([X_orig] if HAS_ORIGINAL else []):
if all(col in df.columns for col in ['Age', 'ST depression']):
df['Age_x_STdepression'] = (df['Age'] * df['ST depression']).astype('float32')
if all(col in df.columns for col in ['Max HR', 'Age']):
df['MaxHR_minus_Age'] = (df['Max HR'] - df['Age']).astype('float32')
for c1, c2, new_col in [('Chest pain type', 'Thallium', 'ChestPain_x_Thallium'),
('Exercise angina', 'Slope of ST', 'ExAngina_x_SlopeST')]:
if c1 in X.columns and c2 in X.columns:
for df in [X, X_test] + ([X_orig] if HAS_ORIGINAL else []):
df[new_col] = (df[c1].astype(str) + '__' + df[c2].astype(str))
Step 3. CV Helper Design
Purpose
Centralize training/evaluation so every model gets exactly the same protocol. This avoids “different training loop, unfair score comparison” problems.
Parameter choices and why
- OOF predictions for unbiased training-set evaluation.
- Test predictions averaged across folds to reduce variance.
- Optional
X_extra/y_extrawithextra_weightto test controlled augmentation. - Early stopping + GPU fallback logic in the full notebook to keep runs practical.
Outcome in this notebook
Produced comparable cv_auc, oof_pred, and test_pred artifacts for all models/variants and all seeds.
Quick notes for beginners:
OOF(Out-Of-Fold) prediction: prediction for each training row made by a model that did not train on that row.- Why OOF matters: it gives a less biased estimate of real generalization than in-fold predictions.
Show key code snippet
Reduced from original notebook: fallback/verbosity and helper details were omitted.
1
2
3
4
5
6
7
8
9
10
11
12
13
14
15
16
17
18
19
20
def run_cv(model_name, model_builder, X, y, X_test, n_splits=5, seed=42,
X_extra=None, y_extra=None, extra_weight=0.0):
skf = StratifiedKFold(n_splits=n_splits, shuffle=True, random_state=seed)
oof = np.zeros(len(X), dtype=float)
test_pred = np.zeros(len(X_test), dtype=float)
for tr_idx, va_idx in skf.split(X, y):
X_tr, y_tr = X.iloc[tr_idx].copy(), y.iloc[tr_idx].copy()
X_va, y_va = X.iloc[va_idx].copy(), y.iloc[va_idx].copy()
if X_extra is not None and y_extra is not None and extra_weight > 0:
X_tr = pd.concat([X_tr, X_extra], axis=0, ignore_index=True)
y_tr = pd.concat([y_tr, y_extra], axis=0, ignore_index=True)
model = model_builder()
model.fit(X_tr, y_tr)
oof[va_idx] = model.predict_proba(X_va)[:, 1]
test_pred += model.predict_proba(X_test)[:, 1] / n_splits
return {'cv_auc': roc_auc_score(y, oof), 'oof_pred': oof, 'test_pred': test_pred}
Step 4. Candidate Models and Tuning Policy
Purpose
Evaluate multiple boosting families under one protocol and optionally tune them with Optuna.
Parameter choices and why
- Enabled candidates:
catboost,lightgbm,xgboost,hgb. PARAM_SOURCE='optuna'in this run:- tune model hyperparameters
- tune
original_weightsimultaneously
- Multiple source modes (
optuna/manual/base) are provided for reuse and reproducibility.
Outcome in this notebook
- Active tuned models loaded into registry:
catboost,lightgbm,xgboost. - Final training proceeded with tuned settings and model-specific original weights.
Quick notes for beginners:
Optuna: automatic hyperparameter search library that tries parameter sets and maximizes an objective (here, OOF AUC).PARAM_SOURCE: switch controlling where final parameters come from (optuna,manual, orbase).model_registry: dictionary of model-builder functions so all models can run under the same training loop.
Show key code snippet
Reduced from original notebook: full parameter grids and trial internals were shortened.
1
2
3
4
5
6
7
8
9
10
11
12
13
14
PARAM_SOURCE = 'optuna'
RUN_TUNING = (PARAM_SOURCE == 'optuna')
TUNING_MODELS = ['catboost', 'lightgbm', 'xgboost']
if PARAM_SOURCE == 'optuna':
ACTIVE_TUNED_PARAMS = dict(TUNED_PARAMS)
ACTIVE_TUNED_ORIGINAL_WEIGHT = dict(TUNED_ORIGINAL_WEIGHT)
model_registry = {
'catboost': lambda: CatBoostClassifier(**cat_params),
'lightgbm': lambda: LGBMClassifier(**lgbm_params),
'xgboost': lambda: XGBClassifier(**xgb_params),
'hgb': lambda: HistGradientBoostingClassifier(**hgb_params),
}
Step 5. Train and Compare: Base vs +Original
Purpose
Test whether external data augmentation improves ranking quality after controlling its influence with weights.
Parameter choices and why
- For each model, run:
base+originalwith model-specific tuned weight
- Use multi-seed CV to make ranking less sensitive to one random split.
- Also evaluate a weighted ensemble from the strongest boosted candidates.
Fold and seed aggregation can be written as:
\[\hat{p}_{\text{test}}(x)=\frac{1}{K}\sum_{k=1}^{K}\hat{p}^{(k)}(x), \quad \hat{p}_{\text{seed}}(x)=\frac{1}{S}\sum_{s=1}^{S}\hat{p}^{(s)}(x)\]and weighted ensemble prediction as:
\[\hat{p}_{\text{ens}}(x)=\sum_{m=1}^{M} w_m \hat{p}_m(x), \quad w_m \ge 0,\ \sum_{m=1}^{M}w_m=1\]Results and interpretation
Leaderboard from notebook output:
| rank | model | variant | cv_auc |
|---|---|---|---|
| 1 | catboost | base | 0.955269 |
| 2 | catboost | +original_w0.7171 | 0.955267 |
| 3 | ensemble | weighted | 0.955236 |
| 4 | lightgbm | base | 0.955166 |
| 5 | lightgbm | +original_w0.4063 | 0.955159 |
| 6 | xgboost | base | 0.955040 |
| 7 | xgboost | +original_w0.3133 | 0.955038 |
| 8 | hgb | +original_w0.3500 | 0.954775 |
| 9 | hgb | base | 0.954771 |
Conclusion for this run:
CatBoost basewas best by a narrow but consistent margin.+originalhelped little and did not surpass top base run.- Ensemble was strong but not top.
Quick notes for beginners:
run_cv_multi_seed: repeats CV with different random seeds and averages predictions for stability.- Ensemble weights satisfy (\sum w_m = 1), so each model contributes proportionally.
Show key code snippet
Reduced from original notebook: optimizer fallback details and print formatting were omitted.
1
2
3
4
5
6
7
8
9
10
11
12
13
14
results = []
for name, builder in model_registry.items():
base_res = run_cv_multi_seed(name, builder, X, y, X_test, SEED_LIST, N_SPLITS)
base_res['variant'] = 'base'
results.append(base_res)
if HAS_ORIGINAL:
w = float(ACTIVE_TUNED_ORIGINAL_WEIGHT.get(name, ORIGINAL_WEIGHT))
aug_res = run_cv_multi_seed(name, builder, X, y, X_test, SEED_LIST, N_SPLITS,
X_extra=X_orig, y_extra=y_orig, extra_weight=w)
aug_res['variant'] = f'+original_w{w:.4f}'
results.append(aug_res)
leaderboard = pd.DataFrame([...]).sort_values('cv_auc', ascending=False)
Step 6. Build Submission
Purpose
Convert the top-ranked run into the competition submission format with no manual ambiguity.
Parameter choices and why
- Select
leaderboard.iloc[0]as the best experiment. - Use that run’s
test_preddirectly for submission.
Outcome in this notebook
- Best model:
catboost - Best variant:
base - Saved file:
./submission.csv - Public leaderboard score from this submission:
0.95334
Quick notes for beginners:
leaderboard.iloc[0]: picks the highest-ranked experiment row after sorting bycv_auc.- Submission format must match competition schema exactly (
id, prediction column name).
Show key code snippet
Reduced from original notebook: print/head formatting was omitted.
1
2
3
4
5
6
7
8
9
best_row = leaderboard.iloc[0]
best_result = next(r for r in results
if r['model_name'] == best_row['model'] and r['variant'] == best_row['variant'])
submission = pd.DataFrame({
ID_COL: test[ID_COL],
TARGET: best_result['test_pred']
})
submission.to_csv('./submission.csv', index=False)
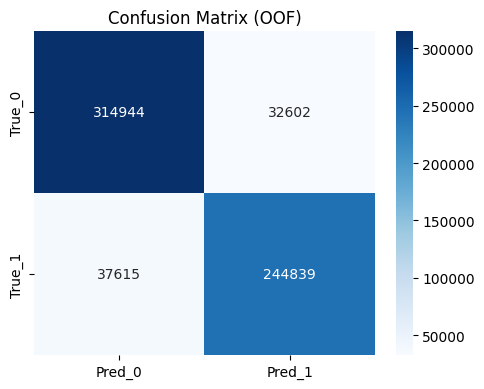
Step 7. OOF Confusion Matrix Check
Purpose
ROC-AUC is threshold-free, but deployment decisions are threshold-based. This section translates probabilistic performance into error-type behavior.
Parameter choices and why
- Threshold fixed at
0.5. - Evaluate confusion matrix and classification report on OOF predictions.
With confusion matrix entries (TP, TN, FP, FN), key metrics are:
\[\mathrm{Accuracy}=\frac{TP+TN}{TP+TN+FP+FN},\quad \mathrm{Precision}=\frac{TP}{TP+FP},\quad \mathrm{Recall}=\frac{TP}{TP+FN}\] \[F_1 = 2\cdot \frac{\mathrm{Precision}\cdot \mathrm{Recall}}{\mathrm{Precision}+\mathrm{Recall}}\]Outcome in this notebook
- TN:
314,944 - FP:
32,602 - FN:
37,615 - TP:
244,839 - Accuracy:
0.8885
Class-wise:
- Class 0: precision
0.8933, recall0.9062, f10.8997 - Class 1: precision
0.8825, recall0.8668, f10.8746
Quick notes for beginners:
confusion_matrix: summarizes classification counts by actual vs predicted class.classification_report: prints precision/recall/F1/support per class in one table.- Threshold (
0.5) is a decision rule; changing it changes FP/FN tradeoff.
Figure 1. OOF confusion matrix at threshold 0.5.
Show key code snippet
Reduced from original notebook: heatmap style arguments were omitted.
1
2
3
4
5
6
7
8
CM_THRESHOLD = 0.5
y_true = y.values
y_prob = best_result['oof_pred']
y_pred = (y_prob >= CM_THRESHOLD).astype(int)
cm = confusion_matrix(y_true, y_pred)
print(classification_report(y_true, y_pred, digits=4))
Supplementary
A. Parameter Source Modes
This notebook supports three parameter-routing modes so you can switch between exploration and reproducible reruns:
optuna: run tuning and apply tuned params/weightsmanual: skip tuning, use manually fixed tuned params/weightsbase: skip tuning, use baseline hyperparameters and globalORIGINAL_WEIGHT
This run used PARAM_SOURCE='optuna'.
B. Detailed Leaderboard (Stability Columns Included)
| model | variant | cv_auc | fold_mean | fold_std | seed_auc_mean | seed_auc_std |
|---|---|---|---|---|---|---|
| catboost | base | 0.955269 | 0.955234 | 0.000004 | 0.955234 | 0.000004 |
| catboost | +original_w0.7171 | 0.955267 | 0.955233 | 0.000004 | 0.955233 | 0.000004 |
| ensemble | weighted | 0.955236 | 0.955236 | 0.000000 | NaN | NaN |
| lightgbm | base | 0.955166 | 0.955032 | 0.000018 | 0.955032 | 0.000018 |
| lightgbm | +original_w0.4063 | 0.955159 | 0.955026 | 0.000013 | 0.955026 | 0.000013 |
| xgboost | base | 0.955040 | 0.954977 | 0.000012 | 0.954977 | 0.000012 |
| xgboost | +original_w0.3133 | 0.955038 | 0.954976 | 0.000011 | 0.954976 | 0.000011 |
| hgb | +original_w0.3500 | 0.954775 | 0.954681 | 0.000018 | 0.954681 | 0.000018 |
| hgb | base | 0.954771 | 0.954676 | 0.000012 | 0.954676 | 0.000012 |
Interpretation:
- Top-2 (
catboost basevscatboost +original) gap is extremely small. seed_auc_stdvalues are low overall, so ranking stability is reasonably strong.
Top-2 gap (absolute and relative):
\[\Delta_{\text{abs}} = 0.955269 - 0.955267 = 0.000002\] \[\Delta_{\text{rel}} = \frac{0.000002}{0.955267}\times 100 \approx 0.000209\%\]This confirms the two CatBoost variants are practically tied in this run.
C. Repro Checklist
If you want to reproduce this exact protocol, keep these fixed:
SEED_LIST = [42, 2024, 7, 3407, 777]N_SPLITS = 5- Same feature preprocessing/category alignment
- Same
PARAM_SOURCEand model enable flags - Same
ORIGINAL_WEIGHTor tuned per-model weights
Final Summary
This notebook is not just “train and submit.” It is a controlled experiment framework with:
- consistent multi-seed CV,
- tunable original-data augmentation,
- optional hyperparameter optimization,
- and explicit model/variant ranking.
For the current run, the answer is obtained by: submit CatBoost (base). Public leaderboard score: 0.95334.